Compensation¶
Compensation adjusts for spectral overlap in your fluorescence data.
Every experiment includes two options for compensation by default. Uncompensated shows the raw data, without any applied compensation. File-Internal is the compensation matrix that was used by the cytometer during data collection. If no compensation was used while collecting data, those matrices will be identical.
Compensation Wizard¶
The compensation wizard uses single-stained controls to automatically generate a compensation matrix. When generating the matrix, both positive and negative peaks are required for each channel. A universal negative control can be used to give a negative peak for every channel, or each single-stained control can include a negative population.
If you don't have single-stained controls, you can edit an existing matrix.
Howto
- In Gating, click on the dropdown underneath compensation.
- Select Create new…. A dialog will open.
- Under Select single-stained controls, check one single-stained sample for each channel.
- Optionally, select a universal negative control.
- Click Next to proceed to channel selection.
- CellEngine will automatically assign FCS files based on which channel has the highest intensity. If needed, assign appropriate files to use for the negative and positive peaks for each fluorescence channel.
- Clean-up channels will be used in the next section to screen out debris or undesired events. CellEngine will default to scatter channels. If other channels are preferred, click on the dropdown beneath Cleanup channels* and choose alternatives.
- Click Next to review gates.
- For each channel:
- If needed, adjust the size and position of the cleanup gate to include the cell or bead population.
- If needed, adjust the positions of the channel gates.
- Click Create, then enter a name to finish generating the matrix.
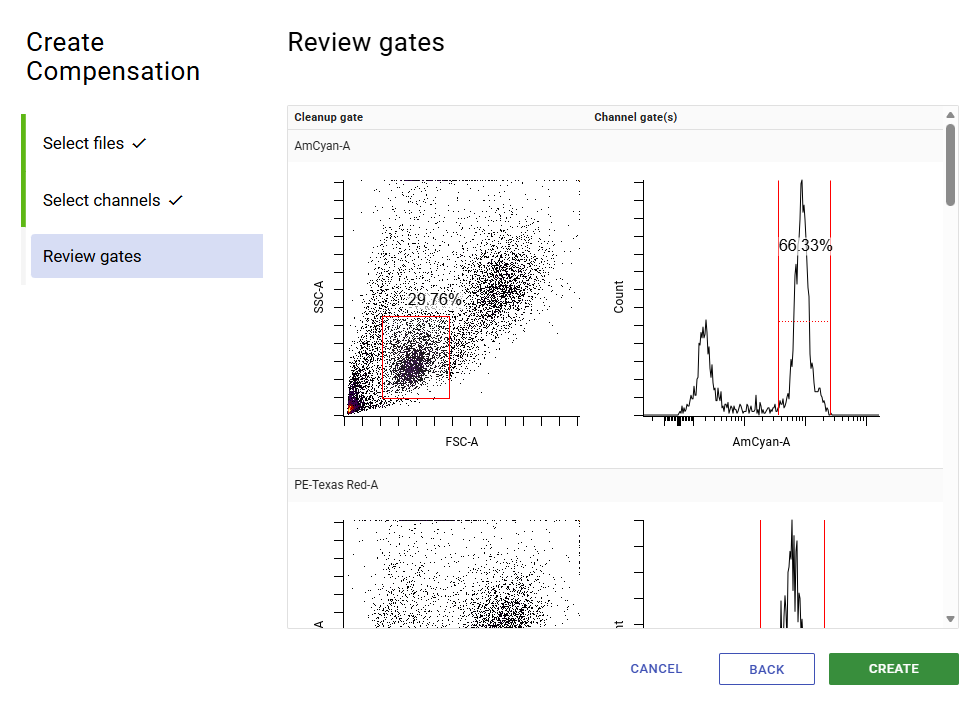
Above is the final step of generating a compensation matrix. Unstained cells were used as the universal negative control, so only the positive gate needs to be set for each channel.
Adjust a Matrix¶
If you want to adjust an existing matrix or create a new matrix without single-stained compensation files, click the Edit button next to the compensation matrix’s name in the gating page to open the compensation editor. You can use your keyboard to adjust compensation values up or down by the increment selected below the matrix. The compensation editor also allows you to adjust your scale settings.
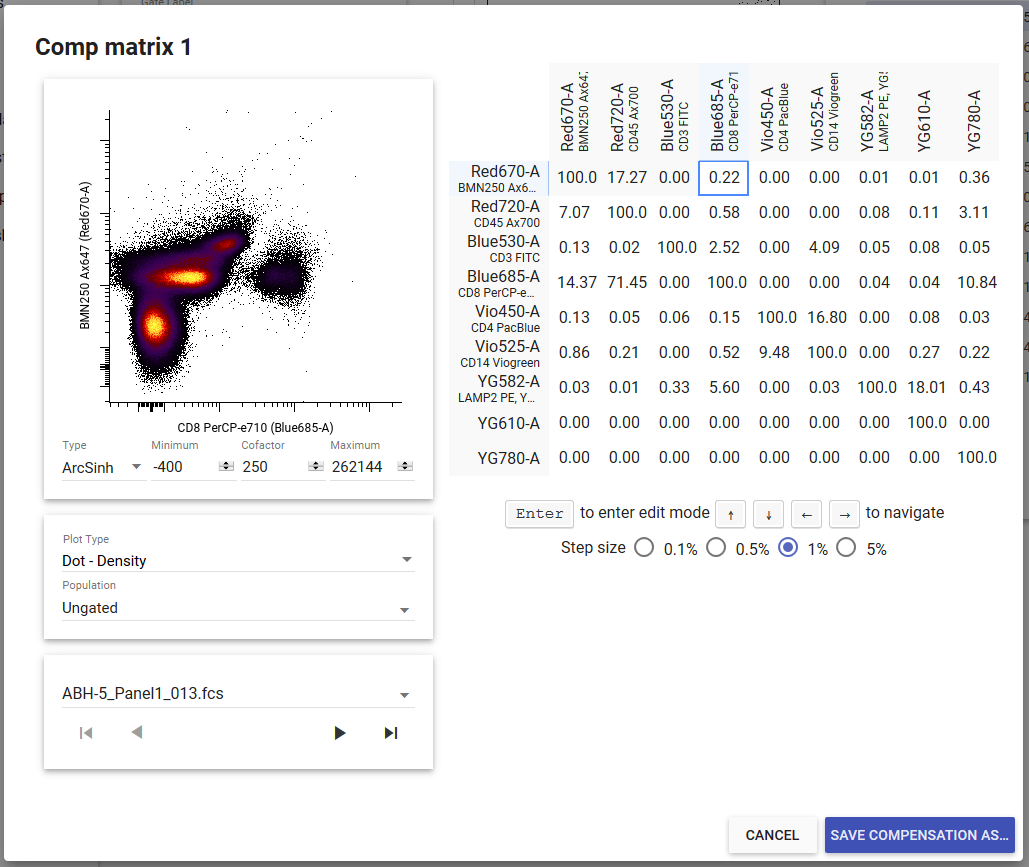
Set Compensation Matrix¶
The compensation matrix can be changed from several places. Changing the compensation matrix on any page will change it for the whole experiment. To change what compensation matrix is used:
Howto
- In Gating, Statistics Export, or Population Export pages, click on the dropdown underneath Compensation, and select the desired matrix.
- In an illustration, click on the matrix name in the toolbar, and select from the dropdown.
Per-File Compensation¶
Warning
This is an advanced feature that, if used inappropriately, will bias your data.
By default, CellEngine applies the same compensation matrix to all FCS files in your experiment. In some cases, you may need to use a different compensation matrix for specific files in the same experiment. For example, if you are running a long-term study and are acquiring data over the course of multiple days, you might have a different compensation on each day due to differences in tandem dye behavior or cytometer performance. You can use per-file compensation in these scenarios.
Howto
- Click Annotations in the sidebar.
- Toggle the per-file compensations option in the toolbar. A new column will appear that lets you select the compensation to use for each file.
- Set your experiment to use per-file compensation from the Gating, Export Populations, Export Statistics, and/or Illustration pages.
Tip
You can use copy/paste keyboard shortcuts to apply a compensation matrix to multiple files at once.
Importing Compensations¶
Compensation matrices can be imported from TSV, CSV, and MTX files exported from CellEngine and FlowJo.
Howto
- From the gating page, click Edit next to the compensation selector. The compensation editor will open.
- In the dotted region in the lower-left corner of the editor, paste a spreadsheet, drag-and-drop a file, or double-click to browse for a file to import.